Lambda phage morphology:
The head has 20 faces. A three-dimensional image with 20 faces is called an icosahedron. The head is made of proteins of various types and contains 46,500 bp long genomic DNA (g). Phage λ contains circular double-stranded DNA approximately 17 µm in length packaged in the protein head of the capsid. The head is 55 nm in diameter and consists of 300 to 600 37,500 Dalton capsomeres (subunits).
Capsomeres are arranged in groups of 5 and 6 subunits, that is, pentamers and hexamers. The head is attached to a 180 µm long non-contractile tail via a connector. The queue consists of 35 stacked disks. It ends in a fibre. There is a hole in the capsid through which this narrow part of the neck passes which expands into a bulge-like structure on the inside. The tail has a thin tail fibre (25 nm long) at its end that recognizes hosts. Also, the tail consists of about 35 stacked discs or rings. Unlike the T-even phage, it is a simple structure devoid of a tail sheath.
Bacteriophage A belongs to the Siphoviridae family of Group I (dsDNA viruses). Lambda Phage Replication Cycle is an E. coli K12 virus that, after entering the host cell, does not normally kill it, despite being capable of destroying the host. Therefore, it carries out its life cycle in two different ways, one as a virulent virus and the second as non-virulent. The virulent phase is called the lytic cycle and the non-virulent as temperate or lysogenic, and the respective viruses as virulent phage and temperate phage, respectively. The other temperate lambdoid phages are 21, Ø80, Ø81, 424, 434, etc.
DNA and gene organization of lambda phage:
Lambda DNA is a linear, double-stranded duplex approximately 17 µm in length. It consists of 48,514 base pairs of known sequences. Both ends of the 5′ terminus consist of 12 bases that extend beyond the nucleotide of the 3′ terminus. This results in a single-stranded complementary region commonly called sticky ends. The sticky ends form base pairs and can easily circularize.
Consequently, a circular DNA with two single-strand breaks is formed. The double-stranded region formed after base pairing of complementary nucleotides is designated COS. The 12 nucleotides at the sticky ends and the process of circularization. Circularization events occur after injection of phage DNA into the E.coli cell where the bacterial enzyme, ie, E.coli DNA ligase, converts the molecule into a covalently sealed circle.
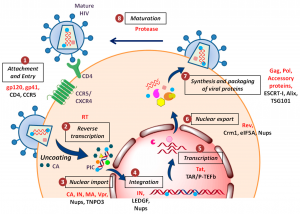
Lambda phage life cycles:
Following adsorption to the host cell’s lamb receptor, lambda gDNA is injected through the tail, which forms a hollow tube through which the DNA passes into the cell. The phage λ goes through two life cycles, the lytic cycle and the lysogenic cycle after injecting its DNA into the E.coli cell. In the lytic cycle, phage genes are expressed and DNA replicates, resulting in the production of various phage particles. The lytic cycle ends with the lysis of E. coli cells and the release of phage particles. This lytic cycle is virulent or moderate in which the phage multiplies into several particles. Furthermore, the lysogenic cycle results in the integration of the phage DNA with the bacterial chromosome and becomes part of the host DNA.
It replicates along with the bacterial chromosome and is inherited in the progeny. The phage DNA integrated with the bacterial chromosome is called a prophage. The prophage is not virulent and is called a moderate phage. Bacteria containing prophage are called lysogenic bacteria and the prophage stage of viruses as lysogenic viruses. After treatment of lysogenic bacteria with ultraviolet light. X-ray or mitomycin, the prophage can separate from the bacterial chromosomes and enter the lytic cycle. This process is known as induction.
Genetic map of phage lambda:
The notable feature of the map is the grouping of genes according to their functions. For example, the head and tail synthesis, replication, and recombination genes are arranged in four distinct groups. These genes can also be grouped into three main operons, viz. right operon, left operon, and immunity operon. The right operon is involved in the vegetative function of the phage, e.g. head synthesis, tail synthesis, and leading lytic cycle in DNA replication.
The left operon is associated with integration and recombination events of the lysogenic cycle. The products of the immunity operon interact with the DNA and decide whether the phage will start the lytic cycle or the lysogenic cycle. Singer et al (1977) have given the nucleotide sequence of ØX174. The genetic map of bacteriophages has been provided by Echols and Murialdo (1978).
(i) Head Synthesis Genes:
On the far left of the phage genome, the major genes, viz. A, W, B, C, D, E are located which are associated with the maturation of phage DNA and head proteins.
(ii) Tail Synthesis Genes:
The F, Z, U, V, G, H, M, L, K, I, J genes are clustered right in the head genes and code for the tail proteins.
(iii) Cleavage and integration genes:
The xis gene encodes the protein that cleaves phage DNA from bacterial chromosomes, and the int-encoded protein is involved in the integration of phage DNA into the bacterial chromosome.
(iv) Recombination:
The two genes int and xis code att P for site-specific recombination. The three red genes code for three proteins at a normal frequency for general recombination. The red is encoded for exonuclease, the red B for beta protein, and the red V for gamma protein. The gamma protein inhibits exonuclease V.
(v) Positive regulation gene:
The N and R genes are the positive regulation genes. The proteins encoded by these genes increase the transcription rate of other genes. The protein encoded by the N gene induces the transcription of the cell, Q, P, A, red, gam, xis, and int, while the protein encoded by the Q gene stimulates the transcription of the head, tail, and lysis genes. The N and Q genes are also required for plaque formation, in the absence of which the number of phage particles would be lower but not zero.
(vi) Negative Regulation Genes:
The cl gene acts as a repressor, and its product maintains the prophage in the lysogenic form in the bacterial host. In addition, cll and cIII help the d gene in lysogeny. Cro-encoded proteins bind to PL and PR and reduce the expression of cl, N, red, and xis genes. Interactions between the cro-encoded Q proteins and the phage repressor occur in the host cell and the result decides the functioning of the lytic or lysogenic cycle. The choice between lysogeny and lysis was discussed in the previous section.
(vii) DNA Synthesis Genes:
The two genes O and P are involved in the synthesis of phage DNA. The origin of DNA replication lies within the coding sequence of the Q gene, which encodes a protein for the initiation of DNA replication, and the gene that generates the sticky ends lies adjacent to one of the ends. The function of the N gene is required in the transcriptional process of these genes.
(viii) Lysis genes:
The S and R genes control the lysis of the bacterial cell envelope that occurs at the end of the lytic cycle.
The choice between lytic and lysogenic cycles:
Shortly after genome circularization and the start of transcription, gpcII and gpcIII accumulate. gpcII binds to PRE (promoter for the establishment of a repressor) and stimulates RNA polymerase binding. gpcIII protects gpcll from degradation by host nucleases.
The lambda repressor (GPL) is rapidly synthesized (B), binds to OL and OR, and inhibits mRNA synthesis and the production of gpcII and gpcIII (proteins) (C). The repressor activates the promoter for repressor maintenance (PRM) which induces the c/ gene to be continuously transcribed at a low rate. This process continues continuously and ensures stable lysogeny when established (C).
Over the course of time, the pro also accumulates. It binds to OL and OR, activates the transcriptional repressor gene cl, and represses PRM function (D). The (gpcl) repressor can block cro transcription. Therefore, there is a race between the production of gpcl and gpcro proteins.